Discovery of PaleRider
PaleRider Information
Morphology: Siphoviridae
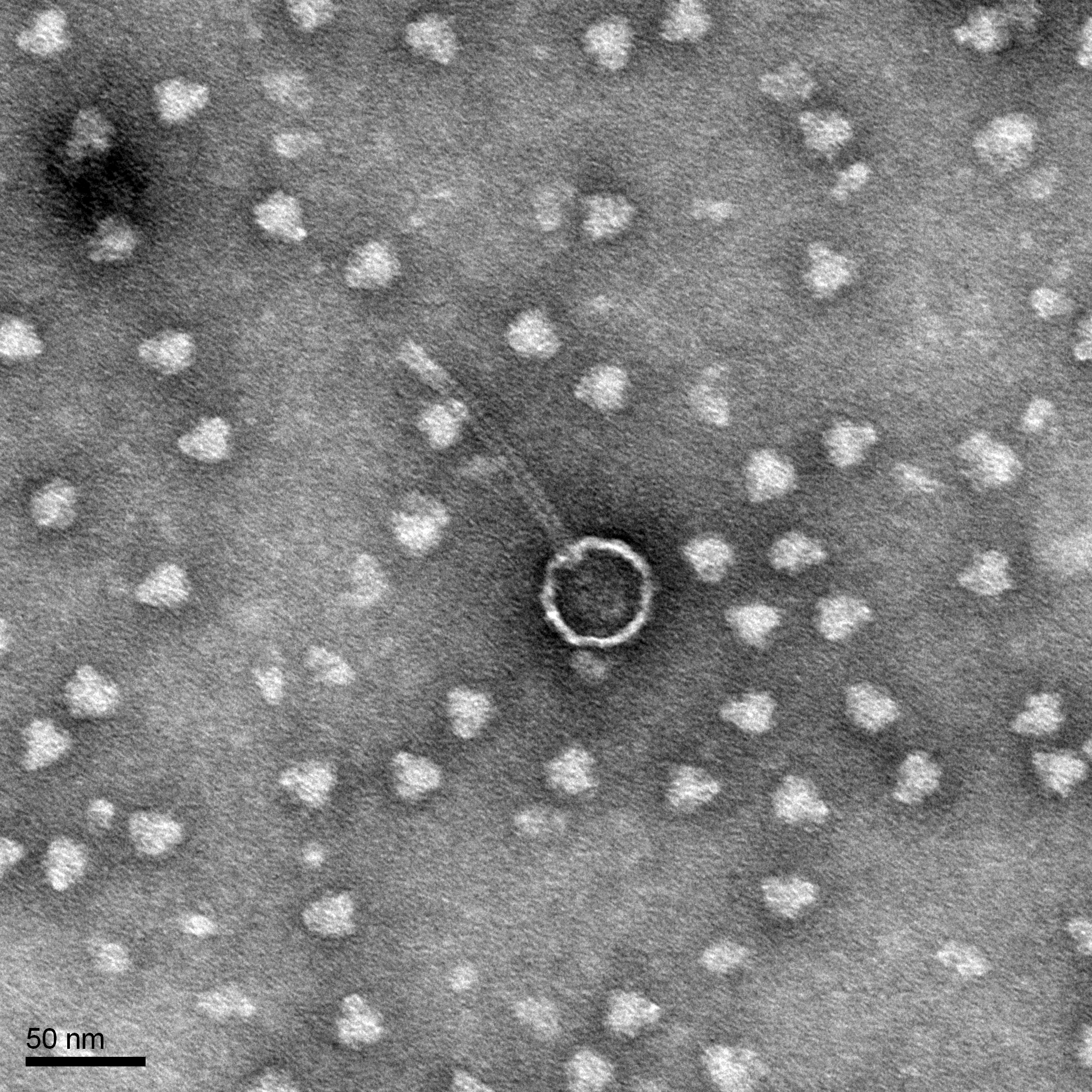
Sample Collection
| Collector Name |
Jacob Schilhab | Jacob Schilhab | Joshua Otero | Joshua Otero | Joshua Otero |
| Sample No. | 1 | 2 | 3 | 4 | 5 |
| Date of Collection | 8/22/2025 | 9/22/25 | 08/22/2025 | 08/22/2025 | 09/22/2025 |
| Sample Type | Soil | Soil | Soil | Soil | Soil |
| General Location | Rising Star, Texas | Stephenville, Texas | Granbury, Texas | Granbury, Texas | Stephenville, Texas |
| Location Description | Underneath shade from trees, near bird dog kennels and a feces pile. | Underneath a tree in a dog park | Hewlett Park, near a small pond with ducks walking around. | Hewlett Park, near a trash can and picnic tables. | Behind a Trash can |
|
GPS Coordinates.
|
32.09707°N, 98.98330°W |
32.44393 °N 98.77866 °W |
32.44325 °N 98.77881 °W | 32.21599 °N 98.22144 °W | |
| Sample Depth | Surface level | Surface Level | Surface Level | Surface Level | Surface level |
| Ambient Temperature | 87°F, 24% Humidity | 75°F, 71% Humidity |
28 ℃, 70% Humidity |
28 ℃, 70% Humidity | 23 ℃, 71% Humidity |
Isolation/Purification
a11
Sample 2 Isolation
Date: 09/23/2025
Redo: Yes
Purpose: The isolation of sample 1 is to determine the presence of bacteriophages for later purification.
Notes:
-
- Extract phages from a soil sample.
- Fill a 50 ml conical tube with your sample to the 15 ml mark.
- Add liquid media to the 35 ml mark and vortex.
- Shake the sample at ~250 rpm for 1–2 hour1.
- Balance the tubes and centrifuge at 2,000 x g for 10 minutes to pellet (i.e., force to the bottom of the tube) most of the soil2.
- Prepare your bench for aseptic work and assemble your supplies.
- Filter the supernatant through a 0.22 µm filter to remove unwanted bacteria and soil particles3.
- Collect the flow-through in a sterile baffled Erlenmeyer flask or a 50 ml sterile conical tube.
- Recovered volumes will range between 20 and 25 ml.
- Filter the supernatant through a 0.22 µm filter to remove unwanted bacteria and soil particles3.
- Seed the culture with host bacteria.
- Add 0.5 ml of bacterial host culture to the flask or conical tube.
- Incubate the flask or conical tube at the proper temperature, shaking at 220 rpm for 2–5 days.
- If you are using a 50 ml conical tube, you must ensure that the culture will be properly aerated. To do so, screw the cap on one-quarter of a turn so that the conical tube is only loosely capped, and then secure the cap with a short piece of lab tape to ensure it does not fall off. Check to make sure that the conical tube remains only loosely capped. Tubes must remain upright while being shaken, and care must be taken to avoid spillage.
- If using a liquid environmental sample, you must add the appropriate volume of 10X liquid media as a source of nutrients for your host bacteria.
- Extract phages from a soil sample.
Results: Enriched isolation was a success, and we were able to proceed into the plaque assay.
Conclusions and Next Steps: We will now proceed into the plaque assay and determine that our plaque contains M. foliorum.
Sample 2 Plaque Assay
Date: 09/29/2025
Redo: Yes
Purpose: The purpose of the plaque assay is to observe the presence of phage particles within our sample visually
Notes: Due to the error of reincubating the plate, our plate dried up, so we had to borrow a fellow group’s plate to proceed to the serial dilution.
-
Day 1
- Prepare your bench for aseptic work and assemble your supplies.
- Assemble the samples you want to assay.
Important: Your phage samples could be taken from your direct isolation, your filtered enriched isolation, or a picked plaque. It is probable that your samples include serial dilutions. - Inoculate the host bacteria with your phage sample.
- Obtain the same number of aliquots of 250 μl host bacterial cultures as you have phage samples, plus one for a positive control and one for a negative control.
- Label the culture tubes accordingly.
- Use a micropipettor and aseptic technique to
- Dispense each phage sample into the appropriate culture tube containing 250 μl of host bacteria according to Table 5.3-1.
- Mix each inoculated host culture by gently tapping the tube.
Important: Make sure your sample makes contact with the bacteria. When you pipette a volume as small as 10 μl sometimes your sample may stick to the side of the tube. - Let your sample sit undisturbed for 5–10 minutes to allow for attachment.
Table 5.3-1. Volumes of phage samples added to bacterial cultures for a plaque assay.
Sample Type
Sample Volume
Direct isolation sample
500 μl
Enriched culture
10 μl
Serial dilutions of picked plaques
10 μl
Lysates for titering
10 μl
Negative control
10 μl phage buffer
Positive control
10 μl provided phage sample
- Obtain the same number of aliquots of 250 μl host bacterial cultures as you have phage samples, plus one for a positive control and one for a negative control.
- Plate the samples with top agar. For this part of the experiment you will need 3 ml of molten top agar per sample. Your instructor may provide this for you or you may need to make it according to the protocol found in the Toolbox.
- Obtain the same number of agar plates as you have samples. (Don’t forget your positive and negative control samples.) Label these plates accordingly.
- Remove a bottle of top agar from the 55°C bath.
- For each sample, including controls
- Using a sterile 5 ml pipette, aseptically transfer 3 ml of top agar to an inoculated host tube (i.e., the tube containing the bacterial host and phage sample).
- Immediately aspirate (suck up) the mixture back into the pipette and transfer it to the appropriate plate, and discard the pipette.
- Gently, but quickly, tilt the plate in multiple directions until the top agar mixture evenly coats the agar plate.
- Repeat this process for each of your samples.
- Incubate plates to allow bacterial growth and phage infection.
- Let the plates sit undisturbed for ~20 minutes until the top agar solidifies.
- After the top agar has solidified, gently invert the plates and place in the proper incubator.
- Incubate the plates at the specified temperature for 24–48 hours.
- Record the incubation time and temperature in your notebook. Results:
Add results and photos here.
Conclusions and Next Steps: The plaque that we had to borrow was sufficient for us to proceed into the serial dilution phase.
Sample 2 Serial Dilution
Date: 10/01/2025
- Prepare your bench for aseptic work and assemble your supplies.
- Set up 10-fold serial dilutions.
- Add 90 μl of phage buffer to each of the tubes.
- Perform 10-fold serial dilutions.
- Add 10 μl of your undiluted phage sample to the “10-1” tube and vortex well.
- The solution in this “10-1” tube contains 1/10th the number of phage particles as your undiluted sample. It is also referred to as a 1:10 dilution.
- Make sure to use a clean pipette tip for each transfer and pipette carefully, vortexing your sample well before making each dilution. Otherwise, you will not make accurate 10-fold dilutions.
- Transfer 10 μl of the “10 -1” sample to the “10-2” tube and vortex well.
- This solution contains 1/100th as many phage particles as your undiluted sample. It can also be referred to as your 1:100 dilution.
- Continue each successive dilution until you get to your last tube.
- Add 10 μl of your undiluted phage sample to the “10-1” tube and vortex well.
Results:
Sample 2: 2nd Serial Dilution
Date: 10/03/2025
- Prepare your bench for aseptic work and assemble your supplies.
- Set up 10-fold serial dilutions.
- Add 90 μl of phage buffer to each of the tubes.
- Perform 10-fold serial dilutions.
- Add 10 μl of your undiluted phage sample to the “10-1” tube and vortex well.
- The solution in this “10-1” tube contains 1/10th the number of phage particles as your undiluted sample. It is also referred to as a 1:10 dilution.
- Make sure to use a clean pipette tip for each transfer and pipette carefully, vortexing your sample well before making each dilution. Otherwise, you will not make accurate 10-fold dilutions.
- Transfer 10 μl of the “10 -1” sample to the “10-2” tube and vortex well.
- This solution contains 1/100th as many phage particles as your undiluted sample. It can also be referred to as your 1:100 dilution.
- Continue each successive dilution until you get to your last tube.
- Add 10 μl of your undiluted phage sample to the “10-1” tube and vortex well.
Results:
Notes: We will be using the 10^-1 dilution for our webbed plate going forward.
Webbed Plate
1. Flooding the 10^-1 plate with phage buffer
2. We are putting the plaque back into the fridge.
10/7/25
3. Lysate was extracted from the webbed plate using a pipette
4. The low-volume lysate was then vacuum filtered using a .22 um filter
5. Approximately 5 mL of low-volume lysate was recovered.
Results: We will continue into phage amplification. using a dilution factor of 10^-2.
Amplification
Making Webbed Plates from a Lysate of Known Titer
Date: 10/8/2025
Redo: No
Purpose: The purpose of this experiment is to create a plate with a very high density of plaques from a lysate of known titer.
Notes: Using the dilution factor of 10^-2, we decided that we need to make 6 plaques to obtain the proper volume.
-
- The procedure for serial dilution was followed to create 6 microcentrifuge tubes, all with a dilution of 10^-2.
- 6 seperate plaques were created with this dilution
- Plates were placed in an incubator for 48 hours.
Results: Upon observation of the plaques after incubation, they were too diluted to continue. We decided to repeat this step using undiluted lysate.
1. 6 microcentrifuged tubes were created without any dilution factors.
2. 6 plaques were placed in the incubator with the undiluted sample.
3. We only allowed the plates to incubate for 24 hours due to their undiluted nature.
Conclusions and Next Steps: The undiluted plates proved successful in creating our webbed plates. The next step will be to collect a high-volume lysate from these webbed plates.
Collecting High-Volume Lysate:
1. 8 mL of sterile phage buffer was added to each webbed plate.
2. The plate was then allowed to sit with the buffer overnight (24 hours) at 4 degrees C.
3. After incubation the buffer was collected with a pipette by putting the plate at an angle.
4. Then after collecting the buffer from the webbed plate, we used a vacuum filter to filter out any agar debris or bacteria in the lysate.
High-volume lysate was collected and stored for future use.
Determining the titer of our high-volume lysate:
In order to continue on with TEM, we need to determine the titer of our high-volume lysate.
1. A ten-fold dilution was created with our lysate with 6 microcentrifuge tubes ranging from 10^-1 to 10^-6.
2. We inoculated the host bacteria, M. foliorum, with the lysate and let it sit for 10 mins.
3. Then, using 3 mL of molten agar, the inoculated bacteria dilutions were transferred to 6 separate plates.
4. Plates were placed in an incubator for…
Characterization
Title
Date: XX/XX/2025
Redo: n/a
Purpose: The purpose of this experiment is to…
Notes:
Results:
Add results and photos here.
Conclusions and Next Steps:
Add conclusions and next steps here.
DNA Extraction
Date: 10/29/2025
Redo: No
Purpose: The purpose of this experiment is to extract and purify phage genomic DNA from high-volume lysate (HVL) using a zinc chloride precipitation method followed by TES lysis, proteinase K treatment, alcohol precipitation, and final purification for downstream Nanodrop quantification.
Notes:
Day 1 – Viral Concentration & Capsid Lysis
-
Mixed HVL gently and aliquoted 5 mL lysate into 15 mL conical tube and added 20 µL nuclease mix, tube inverted gently, incubated 37°C for 10 min.
-
Transferred lysate into five 1.5 mL microfuge tubes (1 mL each).
-
Added 20 µL ZnCl₂ to each tube, inverted to mix, and incubated at 37°C for 5 min to precipitate phage.
-
Centrifuged at 10,000 rpm for 1 min to pellet virions. Pellet expected: small, translucent/whitish.
-
Note pellet visibility: Hard to see
-
-
Removed supernatant carefully, ensuring pellet remained intact.
-
Resuspended each pellet in 500 µL TES buffer and incubated at 60°C for 15 min for capsid denaturation.
-
Added 1 µL Proteinase K and incubated at 37°C for 10 min to remove residual nuclease activity.
-
Added 60 µL potassium acetate, mixed, and placed on ice for 15 min, forming white precipitate (capsid remnants).
-
Centrifuged at 4°C, 12,000 rpm, for 1 min, and collected only the supernatant into new labeled tubes (contains DNA).
-
Added 500 µL isopropanol to each tube, mixed, and placed on ice overnight for DNA precipitation.
Day 2 – DNA Recovery & Purification
-
Centrifuged samples at max speed for 10 min to pellet DNA.
-
Pellet visibility: Faint
-
Washed pellet using 250 µL 70% ethanol, centrifuged 1 min, and discarded supernatant into waste tube.
-
Inverted tubes and air-dried pellets until nearly clear and not visibly wet.
-
DO NOT rush—drying time noted: 45 min.
-
Resuspended the first pellet in 50 µL nuclease-free water and sequentially dissolved all pellets into the same 50 µL final volume.
-
Mixed gently and ensured no visible clumps remained; briefly spun if necessary.
-
Measured DNA concentration and purity using Nanodrop (A260, 260/280, 260/230).
Results:
INSERT PICS OF NANODROP RESULTS HERE
Conclusions and Next Steps:
Next we are going to begin the steps of reprecipitation.